Targeting RNA tertiary structure: discovery of Intronistat B
It is becoming increasingly clear that large, highly structured RNA molecules are essential for most metabolic functions. While therapeutics typically function at the protein level by directly targeting enzymes, receptors and ion channels, small molecules that target specific RNA structures can potentially modulate gene expression; and therefore, protein biosynthesis. This approach represents an entirely new paradigm for drug discovery, providing access to important biological targets previously believed to be undruggable.
Our initial efforts have focused on RNA targets within pathogenic yeasts, as these organisms have become increasingly problematic, particularly for patients with compromised immune systems, such as recipients of implanted devices, neonatal patients, and cancer patients 1. The availability of potent antifungals that lack toxicity in mammals is a major unmet medical need, and of value for industrial and agricultural applications. The development of new antifungals is difficult because, as eukaryotes, fungi and yeast cells have enzymes and biochemical pathways that are similar to those of humans. However, fungal RNA metabolism differs significantly, thereby providing a potential route toward new therapeutics 2, 3.
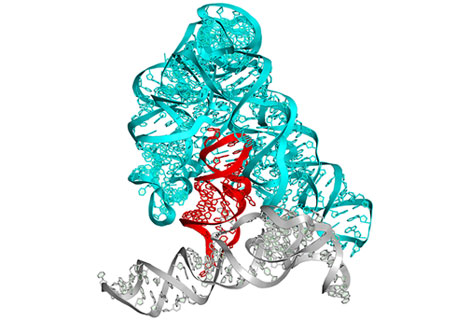
To meet these challenges, we set out to identify small molecule inhibitors of group II introns, which are large self-splicing ribozymes found in the mitochondrial genomes of plants, fungi, and yeast, but are not present in mammals. These autocatalytic RNA molecules adopt an elaborate tertiary structure that has been crystallographically characterized and which contains an active-site for RNA cleavage and ligation as well as solvent-accessible pockets for potential inhibitor binding 4, 5 (Fig. 1a).
In yeasts such as Saccharomyces cerevisiae (S. cerevisiae) and the pathogen Candida parapsilosis (C. parapsilosis), group II introns are found within genes that are essential for respiration, such as cytochrome oxidase subunit genes of the mitochondria 6,7. Importantly, respiration is essential for pathogenic yeast to differentiate into biofilms, which colonize medical implant surfaces, are resistant to antifungals, and contribute to pathogenic virulence 2, 3, 7. Based on the complexity of their structures, and their essential role in fungal metabolism, group II introns represent outstanding targets for the development of highly specific antifungal agents.
Figure 1.
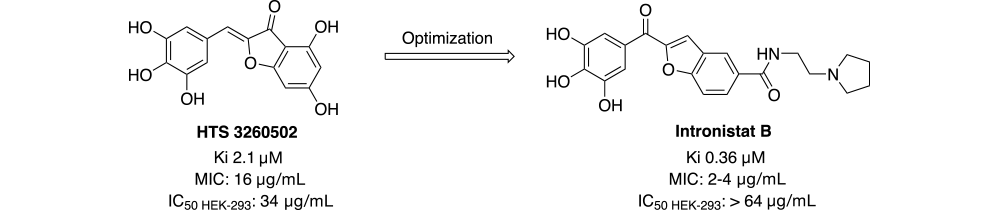
Using an HTS assay developed to monitor yeast group II intron splicing reactivity, several promising hits were identified including, HTS-3260502, which has a Ki of 2.1 µM and inhibited fungal growth with a minimal inhibitory concentration (MIC) of 16 µg/ml. Optimization of this early lead led to a series of compounds now known as Intronistats. Intronistat B, the most potent compound in the series has a Ki of 0.36 µM and MIC of 2-4 µg/mL. In addition to increasing the potency in both the screening and fungal growth assays by approximately 6-fold relative to HTS-3260502, Intronistat B also was superior in a cell viability assay (HEK293), where no effect was observed at 64 µg/mL, the highest concentration tested.
Compounds in the series were prepared from readily available and inexpensive 3, 4, 5-trimethoxyacetophenone. Treatment with isopropenyl acetate with tosic acid followed by NBS gives bromomethyl ketone 3 in 90% yield (2 steps). Subsequent condensation with methyl 3-formyl-4-hydroxybenzoate 4 using potassium carbonate in MEK gives ketone 5. Hydrolysis of the methyl ester with aqueous sodium hydroxide in methanol followed by EDC coupling with amine 7 gives the penultimate intermediate 8. Deprotection on the methoxy substituents using boron tribromide gives Intronistat B in six steps with an overall yield of approximately 30%.
Scheme 1.
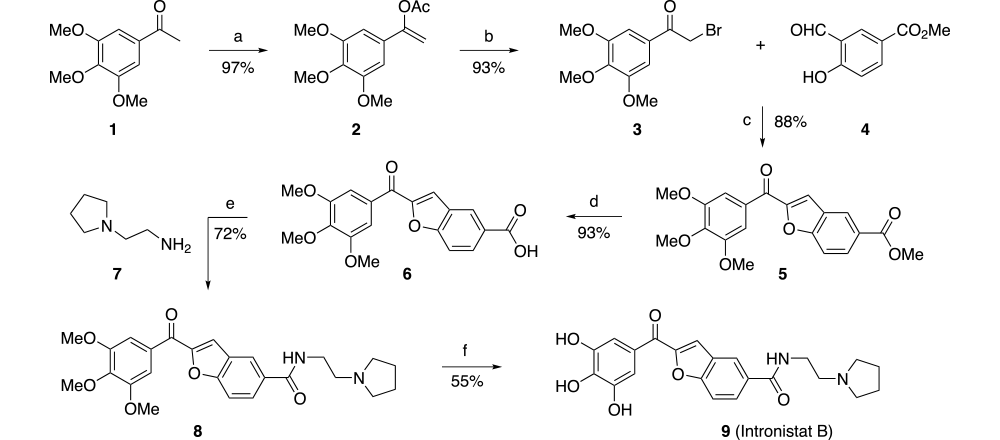
Reagents and conditions: (a) isopropenyl acetate, TsOH, reflux; (b) NBS, THF; (c) K2CO3,MEK 50 ºC,; (d) Aq NaOH, MeOH; (e) HOBt, EDC, CH2CI2; (f) BBr3, CH2CI2.
These studies demonstrate that biologically important RNA molecules can be targeted with high specificity, potency and efficacy using the tools of medicinal chemistry and microbiology, resulting in a promising new class of antimicrobials that are likely to be far less toxic to humans than existing types of therapy. They also demonstrate that structured RNA molecules can be targeted de-novo using the same approaches that have upheld the pharmaceutical industry for the last century.
This important work, which was published in Nature Chemical Biology (Fedorova et al, Nature Chemical Biology (2018) 14, 1073-1078), has been the subject of commentary in C&E News, Nature Reviews Drug Discovery, Gen&En News and other media outlets (see links below).
Fedorova et al, Nature Chem. Biol. (2018)
www.ncbi.nlm.nih.gov/pubmed/30323219
Palacino, “The splice is right.” J. News and Views in Nature Chem. Biol. (2018)
www.ncbi.nlm.nih.gov/pubmed/30323218
Nature Reviews Drug Discovery, “Small molecules targeting a tertiary RNA structure fight fungi.”
www.ncbi.nlm.nih.gov/pubmed/30482965
GenEngNews: “Fungal RNAs targeted.”
www.genengnews.com/news/fungal-rnas-targeted-by-small-molecule-drugs/
Chem&EngNews: “Targeting RNA’s tertiary structure.”
cen.acs.org/pharmaceuticals/drug-discovery/Targeting-RNAs-tertiary-structure/96/i43
References
- Kett, D. H., Azoulay, E., Echeverria, P. M., Vincent, J. L. & Extended Prevalence of Infection in, I. C. U. S. G. o. I. Candida bloodstream infections in intensive care units: analysis of the extended prevalence of infection in intensive care unit study. Crit Care Med 39, 665-670, doi:10.1097/CCM.0b013e318206c1ca (2011).
- Richard, M. L., Nobile, C. J., Bruno, V. M. & Mitchell, A. P. Candida albicans biofilm-defective mutants. Eukaryot Cell 4, 1493-1502, doi:10.1128/EC.4.8.1493-1502.2005 (2005).
- Morales, D. K. et al. Control of Candida albicans metabolism and biofilm formation by Pseudomonas aeruginosa phenazines. MBio 4, e00526-00512, doi:10.1128/mBio.00526-12 (2013).
- Toor, N., Keating, K. S., Taylor, S. D. & Pyle, A. M. Crystal structure of a self-spliced group II intron. Science 320, 77-82, doi:10.1126/science.1153803 (2008).
- Toor, N. et al. Tertiary architecture of the Oceanobacillus iheyensis group II intron. RNA 16, 57-69, doi:10.1261/rna.1844010 (2010).
- Askew, C. et al. Transcriptional regulation of carbohydrate metabolism in the human pathogen Candida albicans. PLoS Pathog 5, e1000612, doi:10.1371/journal.ppat.1000612 (2009).
- Rossignol, T. et al. Correlation between biofilm formation and the hypoxic response in Candida parapsilosis. Eukaryot Cell 8, 550-559, doi:10.1128/EC.00350-08 (2009).

